Spatial DNA Methylation
Available in 2026
Map DNA methylation in tissue, with spatial precision
A next-generation assay that localizes methylation patterns directly in tissue sections—enabling multi-omic context, cell-type resolution, and disease-relevant insights you can’t get from bulk or dissociated methods.
Single-slide workflow
Tissue-native context
Multi-omics compatible
Now inviting collaborators for early access evaluation.
AGBT 2026 Poster
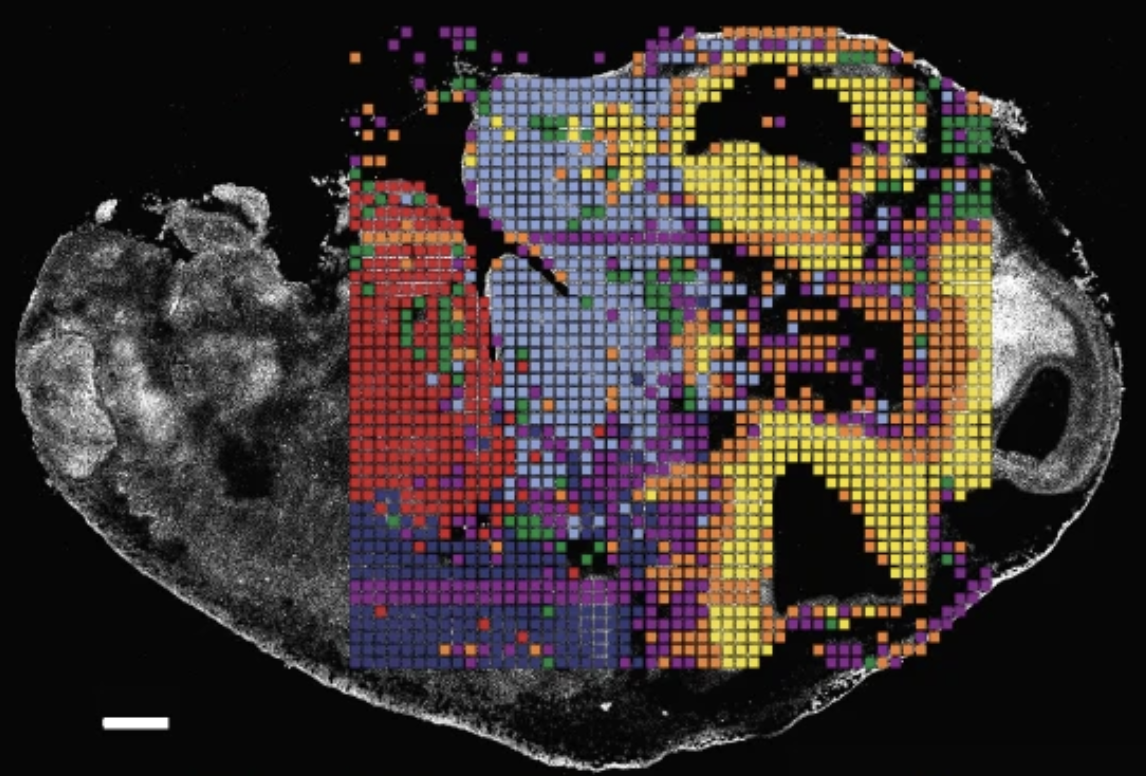
Spatial ATAC-TAPS+ integrates chromatin accessibility and DNA methylation profiling in a single tissue section, resolving epigenetic heterogeneity that bulk methods miss.
New Research
Spatial ATAC-TAPS+: Co-Registered Accessibility & Methylation Profiling
- 27,923 clustered nuclei at 10 µm resolution
- 8 distinct epigenetic compartments in human medulloblastoma
- 2,195 differentially methylated genes identified
- TAPS+ conversion preserves ATAC library quality

Join Early Access
Get in touch


Email Us:
Info@atlasxomics.com
Vist Us:
C/O John B. Pierce Laboratory
290 Congress Ave.
New Haven, CT 06519-1403
Follow our research

