Map Gene Regulation and Expression with Spatial Context
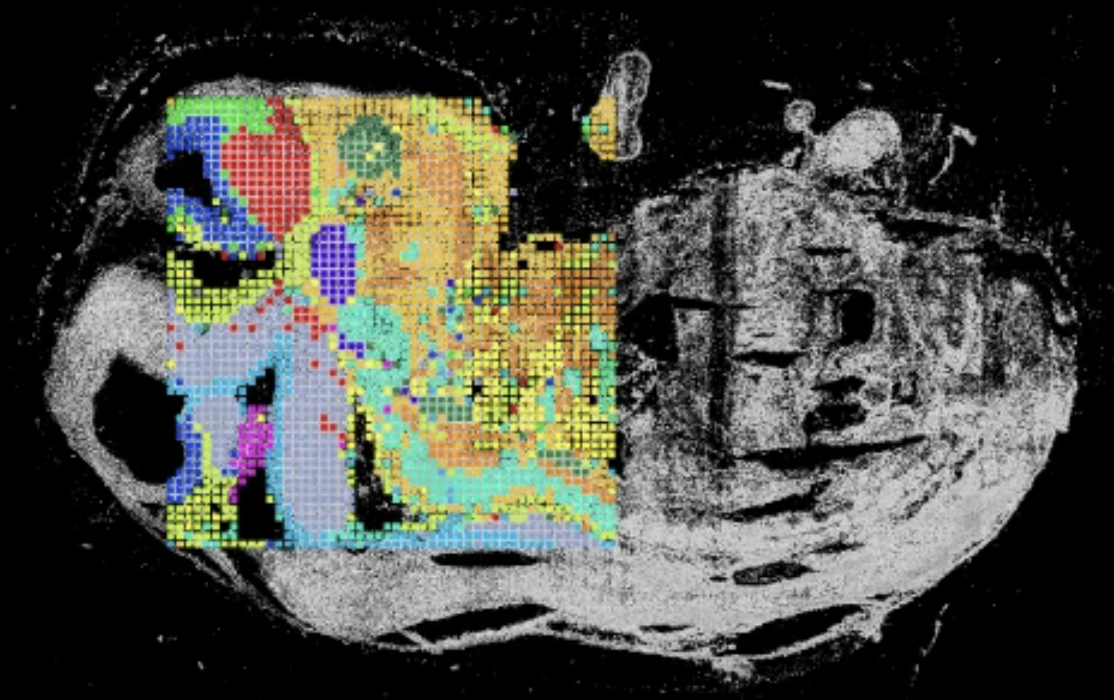
Spatial CoProfiling simultaneously measures chromatin accessibility and RNA from the same tissue section—directly linking regulatory state to transcriptional output, pixel by pixel, within intact tissue architecture.
The assay is built on spatial ATAC–RNA-seq, first described in a 2023 Nature publication from the Fan Lab at Yale, which demonstrated near-single-cell resolution co-profiling of the epigenome and transcriptome in mammalian tissues.
Featured Resources
Explore the white paper and conference poster to see CoProfiling applied across tissue contexts.
Spatial CoProfiling (ATAC + RNA)
- Paired ATAC + RNA from the same section
- Direct regulatory interpretation
- Spatially resolved chromatin–expression coupling

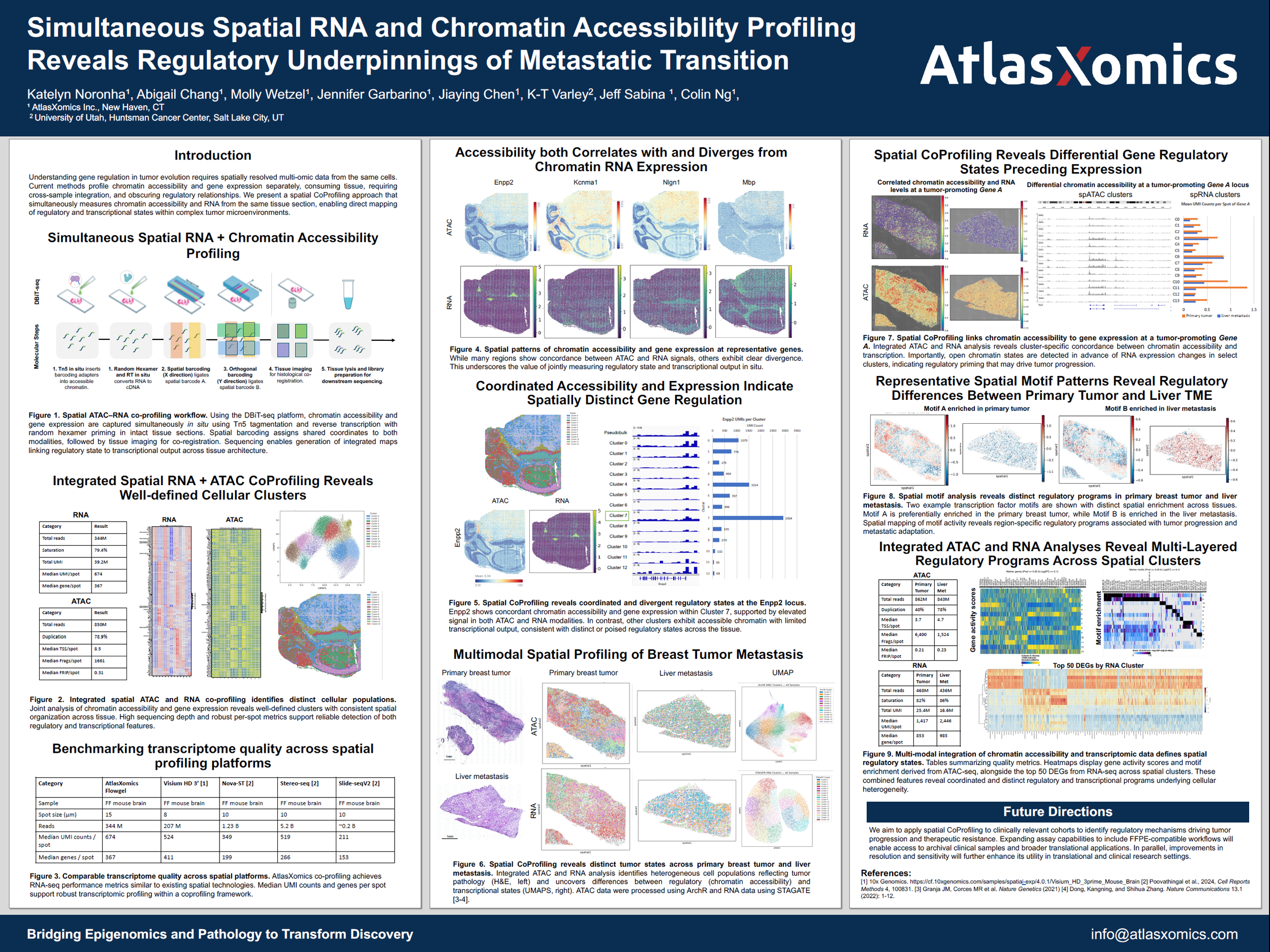
Simultaneous Spatial RNA and Chromatin Accessibility Profiling Reveals Regulatory Underpinnings of Metastatic Transition
- Open chromatin detected ahead of RNA changes, revealing regulatory priming in select tumor clusters
- Distinct transcription factor motif enrichment between primary breast tumor and liver metastasis
- Integrated ATAC + RNA clustering identifies heterogeneous tumor states across tissue architecture
Join Early Access
Get in touch


Email Us:
Info@atlasxomics.com
Vist Us:
C/O John B. Pierce Laboratory
290 Congress Ave.
New Haven, CT 06519-1403
Follow our research
